We pioneered the machine learning and predictions of chromatographic behavior (Palmblad et al. 2002) and use of isotopic distributions in proteomics (Palmblad et al. 2001) - principles now widely adopted in proteomics. We have particular expertise in multiplexing schemes for quantitative proteomics (Palmblad et al. 2008), integrating large proteomics datasets, and published the first population-scale proteomics in 2013 (Johansson et al.).
Other work include bridging gaps between human and model system proteomics, in particular zebrafish (van der Plas-Duivesteijn et al. 2014) and mouse (Mohammed et al. 2021), proteomics in non-model systems such as pathogens or parasites (Yilmaz et al. 2016) and contextualizing omics data combining text mining and computational chemistry (Palmblad 2019). Ongoing work include data refinement and reanalysis (Marissen and Palmblad 2021), improved methods for and novel applications of direct comparison of tandem mass spectra (Marissen et al. 2022) and looking to the future by investigating computational aspects of next-generation proteomics (Palmblad 2021).
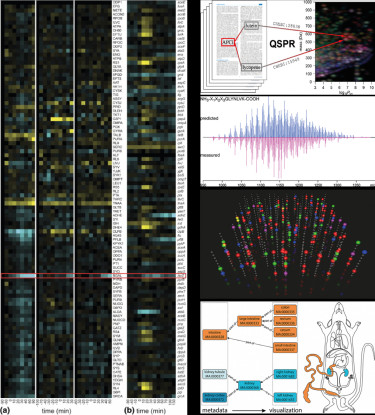
Collage of recent work from the Bionformatics Group.


